Global health officials struggling to curb the spread of malaria continue to face the additional challenge of drug resistance — leaving scientists in search of new models for effective treatment and prevention.
In response to this problem, a team at the University of Notre Dame put out a call to the masses, enlisting researchers, data scientists and health professionals to analyze genomic data from emerging drug-resistant malaria parasites and gain a better understanding of the mechanisms of resistance.
Three hundred and sixty participants from 31 countries participated in the Malaria DREAM (Dialogue for Reverse Engineering Assessments and Methods) Challenge, a crowdsourcing effort challenging anyone in the world to develop computational models for predicting emerging drug resistance to artemisinin, a widely used therapeutic considered the “last line of defense” against multi-drug-resistant malaria.
 Geoffrey Siwo
Geoffrey Siwo
This is the first DREAM challenge to address an infectious disease that predominantly affects the developing world.
“Crowdsourcing has a long history of developing new and improved solutions for big problems by posing those problems to communities around the world,” said Geoffrey Siwo, research assistant professor in the Department of Biological Sciences and the Eck Institute for Global Health at Notre Dame, who led the challenge. “While the method has been used to address issues in cancer research, no DREAM Challenge had looked at issues related to infectious diseases like malaria. Malaria receives less funding than cancer, there are fewer technologies available to address the problem of infection and, now, drug resistance, and it can take nearly 10 years before new machine learning methods are applied to the disease.”
DREAM Challenges, a nonprofit organization, has attracted researchers from universities, technology companies and biotechnology and pharmaceutical companies. Siwo, along with Michael Ferdig, also a professor in the Department of Biological Sciences at Notre Dame, and DREAM Challenges organized the Malaria DREAM Challenge.
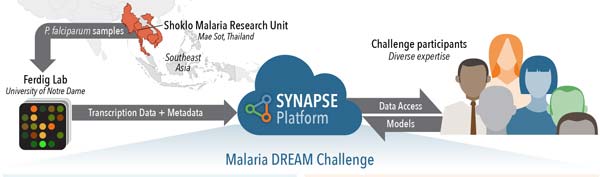
The emerging artemisinin resistance is difficult to measure both in malaria-infected patients and using laboratory experiments. So, the challenge asked participants to determine how genomic data of parasites can be used to predict their drug resistance levels based on patient-derived data and laboratory experiments. In their paper, “Leveraging crowdsourcing to accelerate global health solutions,” published in Nature Biotechnology, the challenge organizers provided an overview of the challenge to the scientific community. The challenge opened to participants in April and closed in September.
Two teams tied for first place in modeling the levels of dug resistance using a measure of resistance obtained in laboratory experiments, known as inhibitory concentration 50 or IC50. The teams were led by Jinseub Hwang from Daegu University in South Korea and Slim Fourati from Case Western Reserve University. The top team in the modeling of the resistance levels using another measure of drug response directly obtained from patients, commonly referred to as parasite clearance rate, was led by Yuanfang Guan from University of Michigan.
The best performing teams presented their work at the Regulatory and Systems Genomics with DREAM challenges meeting held from November 4 to 6th 2019 at the Rockefeller Research Laboratories in New York.
“The challenge has demonstrated that with the right modeling approaches, predicting artemisinin drug response of malaria parasites using genomic data is a tractable problem,” Siwo said. “More than 300 modelers were able to work on this single problem for four months, and through the source code and write-ups they have submitted in this challenge, they have contributed an enormous amount of knowledge that would take years to accrue. When biomedical problems are posed to the crowd, you have more eyes on a singular problem, in a way that even the largest company in the world cannot afford.”
The collaboration brought together several international organizations including the National Institutes of Health, Sage Bionetworks, IBM Research, Texas Biomedical Research Institute, Grand Valley State University, Icahn School of Medicine at Mount Sinai, H3Africa Bioinformatics Network, University of Cape Town, Institut Pasteur de Tunis, Abdelmalek Essaadi University and the Mahidol Oxford Tropical Medicine Research Unit.
The National Institutes of Health and the Bill & Melinda Gates Foundation funded the challenge.
Siwo is affiliated with Notre Dame’s Center for Research Computing.
Contact: Jessica Sieff, assistant director of media relations, 574-631-3933, jsieff@nd.edu