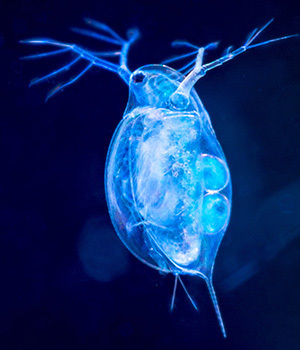
For many, experience with Daphnia, commonly known as water fleas, ends in high school. The organism is often used for science experiments exploring water toxicity, because of its sensitivity to environmental factors. But the tiny, transparent microcrustaceans have been studied intensively for more than 150 years, and new research published and featured on the cover of the journal G3 reveals scientists can now take a closer look at its genome.
Researchers have completed a new and improved genome sequence of Daphnia pulex (D. pulex), providing a clearer roadmap of the organism’s genome so they can identify the genes and pathways that make this organism so successful in freshwater ecosystems.
Populations of Daphnia, barely visible to the naked eye, can be found in virtually every standing body of water on the planet, including Antarctica. They evolve quickly and are masters of responding to the conditions in their environment. Sensing the chemical cues of nearby predators, some species of Daphnia develop elaborate defensive structures such as spines and helmets that make them harder to eat. While scientists have gained a thorough understanding of what these tiny water fleas do to adapt to varying conditions, they don’t yet know how they do it.

“That’s why a system like this is so powerful,” said Michael E. Pfrender, director of the Genomics & Bioinformatics Core Facility and associate professor in the Department of Biological Sciences and the Environmental Change Initiative at the University of Notre Dame. “We need this genomic infrastructure to add to the ecological context we already have to gain a better understanding of how Daphnia adapt. Because we have an improved genome sequence, we can get a more accurate catalog of genes and when thinking about response to the environment and chemical cues, it’s the turning on and off of genes and pathways that’s important. The picture is much more complete than it was before.”
Calling it the “Portland Arch” genome after the Indiana Nature Preserve where the Daphnia was collected, the new assembly comes six years after the first sequence of D. pulex in 2011. The current study describes how scientists used the latest technology as part of a thorough and methodical process the result of which led to the identification of 18,440 genes.
D. pulex plays a vital role in Earth’s ecology. Feeding off of algae and phytoplankton in standing freshwaters, they are the primary grazer in those environments, the “cows of lakes,” said Pfrender. They’re also primary forage, transferring all of that energy to the fish that eat them. By understanding how species of Daphnia respond to toxic elements like industrial contaminants, toxic algae blooms or thermal stress, scientists can look at how environmental changes caused by agriculture and road runoff or warming temperatures and climate change could impact populations in lakes, rivers and standing bodies of water.
“What happens to this vital part of the ecosystem when conditions change very rapidly? What genes allow some populations to cope with these changes while others fail?” Pfrender said. “That’s what we want to find out. This genome sequence provides the toolkit.”
Co-authors of the study include Pfrender, Jacqueline Lopez and Brent Harker of the Notre Dame Genomics and Bioinformatics Core Facility; Zhiqiang Ye, Ken Spitze, Xiaoqian Jiang, Matthew S. Ackerman and Michael Lynch at the Department of Biology at Indiana University; Sen Xu at the Department of Biology at Indiana University and the University of Texas at Arlington; Jana Asselman, Laboratory for Environmental Toxicology at Ghent University; R. Taylor Raborn at the Department of Biology and School of Informatics and Computing at Indiana University; and W. Kelley Thomas and Jordan Ramsdell at the Hubbard Center for Genome Studies at the University of New Hampshire.
The study was funded through a grant from the National Institutes of Health to Michael Lynch at Indiana University, Bloomington and Notre Dame Research.
Contact: Michael Pfrender, 574-631-0591, michael.pfrender.1@nd.edu