University of Notre Dame biologist Michael Pfrender is the coauthor of a paper appearing today in the prestigious journal Science describing the sequencing of the species Daphnia pulex, often referred to as the water flea. Daphnia, a small freshwater crustacean, is the first crustacean to have its genome sequenced. It contains more than 31,000 genes, as compared to 23,000 genes of humans.
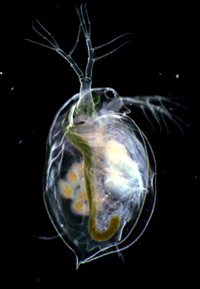
“ Daphnia has long been a model for ecology and early in the last century significant biological findings had their origin in Daphnia ,” Pfrender said. “For example, the concept of phenotypic plasticity, the dose-response curve central to toxicology and the effects of inbreeding to name a few.”
Pfrender was one of a small group of scientists who first approached the U.S. Department of Energy’s Joint Genome Institute in 2002 with a suggestion to sequence the Daphnia genome. That suggestion evolved into the Daphnia Genomics Consortium , an international group of some 450 scientists from dozens of institutions world-wide.
The consortium scientists were interested in the species because of its importance in aquatic food webs and for its transformational responses to environmental stress. Some species of Daphnia produce exaggerated spines, neck-teeth or helmets as a means of self-defense from predators. Daphnia also thrives in the absence of males, using clonal reproduction, until harsh environmental conditions favor the benefits of sex.

Although Pfrender and the founding members of the consortium conceived of the project with the expectation that new gene functions would be uncovered when studied in light of the animal’s natural environment, they did not expect to discover so many more genes.
The researchers reveal in their Science paper that the high number stems from the fact that Daphnia is creating copies of genes at an extremely high rate: more than three times that of other invertebrates and 30 percent greater than that of humans.
“These duplicate genes rapidly acquire new functions in response to environmental conditions,” Pfrender said. “This evolutionary flexibility may, in part, explain why Daphnia is able to respond to the environment in such a flexible manner.”

The genome sequencing information described in Science can help lead to a better understanding of what genes matter for organisms to cope with environmental stress that includes pollutants and global warming. This project used modern functional genomic technologies to understand how these genes function within an animal that is easily studied in fresh water ecosystems around the globe.
Pfrender notes that the collaborative sequencing effort has positioned Daphnia as a model organism for an emerging scientific field of environmental genomics, aimed at better understanding how the environment and the genes in natural populations interact.
“As a genetic model, Daphnia was for some time supplanted by organisms like Drosophila ,” Pfrender said. “ Daphnia is now poised to become the premier model for ecological and environmental genomics. This paper establishes the foundation for the next wave of research in these fields.”
The Daphnia sequencing research received financial and material support from the Office of Science of the U.S. Department of Energy, the National Science Foundation, the Lilly Endowment Inc., Roche NimbleGen Inc., the National Institutes of Health, the U.S. Department of Health and Human Services and Indiana University.